Biochemistry
|
11 november 2018 02:00:14 |
| The structure of the deubiquitinase USP15 reveals a misaligned catalytic triad and an open ubiquitin-binding channel [Enzymology] (Journal of Biological Chemistry) |
|
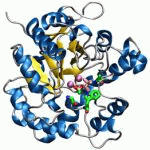
Tweet Ubiquitin-specific protease 15 (USP15) regulates important cellular processes, including transforming growth factor ? (TGF-?) signaling, mitophagy, mRNA processing, and innate immune responses; however, structural information on USP15`s catalytic domain is currently unavailable. Here, we determined crystal structures of the USP15 catalytic core domain, revealing a canonical USP fold, including a finger, palm, and thumb region. Unlike for the structure of paralog USP4, the catalytic triad is in an inactive configuration with the catalytic cysteine ~10 Å apart from the catalytic histidine. This conformation is atypical, and a similar misaligned catalytic triad has so far been observed only for USP7, although USP15 and USP7 are differently regulated. Moreover, we found that the active-site loops are flexible, resulting in a largely open ubiquitin tail-binding channel. Comparison of the USP15 and USP4 structures points to a possible activation mechanism. Sequence differences between these two USPs mainly map to the S1´ region likely to confer specificity, whereas the S1 ubiquitin-binding pocket is highly conserved. Isothermal titration calorimetry monoubiquitin- and linear diubiquitin-binding experiments showed significant differences in their thermodynamic profiles, with USP15 displaying a lower affinity for monoubiquitin than USP4. Moreover, we report that USP15 is weakly inhibited by the antineoplastic agent mitoxantrone in vitro. A USP15-mitoxantrone complex structure disclosed that the anthracenedione interacts with the S1´ binding site. Our results reveal first insights into USP15`s catalytic domain structure, conformational changes, differences between paralogs, and small-molecule interactions and establish a framework for cellular probe and inhibitor development. |
| 98 viewsCategory: Biochemistry |
 Structural and spectroscopic analyses of the sporulation killing factor biosynthetic enzyme SkfB, a bacterial AdoMet radical sactisynthase [Protein Structure and Folding] (Journal of Biological Chemistry) Structural and spectroscopic analyses of the sporulation killing factor biosynthetic enzyme SkfB, a bacterial AdoMet radical sactisynthase [Protein Structure and Folding] (Journal of Biological Chemistry)Identification, functional characterization, and crystal structure determination of bacterial levoglucosan dehydrogenase [Protein Structure and Folding] (Journal of Biological Chemistry) 
|
| blog comments powered by Disqus |
MyJournals.org
The latest issues of all your favorite science journals on one page
The latest issues of all your favorite science journals on one page